Human GRCh37 patch release 8
contains an update to previously released fix patch HG1211_PATCH.
This encompasses a 3Mb region in
GRCh37 between chr10: 46,256,855-49,299,273.
The tile path in the 10q11.22 region
has been extensively altered from its previously fragmented state to one where
a single gap remains, between BX649215.1 and AC245041.3. The reworking of the
tile path in the region has been carried out using clones in the existing build
and additional finished clones not previously in GRCh37.
Working with optical map data
provided by the Schwartz Lab we have been able to identify errors in the GRCh37
assembly and have consequentially worked to correct them. The optical map
analysis also highlighted redundancy in the assembly causing artificial
duplication, which has now been addressed within this patch.
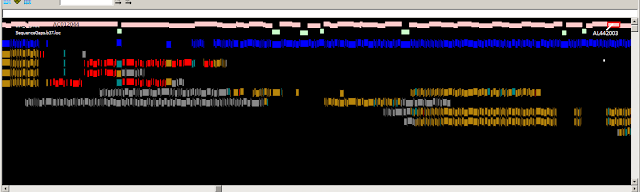 |
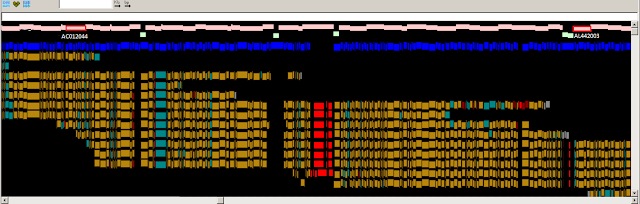
Above: Optical map consensus alignments to GRCh37 10q11.22.
Below: Optical map consensus alignments to the fix patch (JH591181.2)
Legend: Pink track: Clone path; Green: Contig gap; Blue: In silico SwaI fragments.
For the aligned optical map consensus Gold: Concordant fragment ; Red: Missing fragment (seen where OM consensus span gap); Grey: Unaligned fragment
|
The optical map information was consistent with a path problem in this region. The map data suggested that several clones in the region were misplaced and did not represent a valid chromosome structure in this region. In addition to rearranging several clones (including changing the orientation of some clones in the path), 3 finished clones were added to the path and several redundant clones were removed. The new path contains a single gap that we estimate, based on optical mapping, to be about 90 Kb. The figure below shows an alignment of the patch sequence to the current chr10 assembly.

I was diagnosed as HEPATITIS B carrier in 2013 with fibrosis of the
ReplyDeleteliver already present. I started on antiviral medications which
reduced the viral load initially. After a couple of years the virus
became resistant. I started on HEPATITIS B Herbal treatment from
ULTIMATE LIFE CLINIC (www.ultimatelifeclinic.com) in March, 2020. Their
treatment totally reversed the virus. I did another blood test after
the 6 months long treatment and tested negative to the virus. Amazing
treatment! This treatment is a breakthrough for all HBV carriers.