The Rat Genome Database RGD is very pleased to announce the release of mRatBN7.1, the new rat genome assembly! The
mRatBN7 assembly, generated by the Darwin Tree of Life Project at the Wellcome Sanger Institute, is significantly improved over the Rnor6.0 and
previous assemblies. mRatBN7 was derived from a male BN/NHsdMcwi rat that is a direct descendent from the female BN rat previously sequenced. The new BN rat reference genome was generated using multiple technologies including PacBio long reads, 10X linked reads, Bionano maps and Arima Hi-C. Its quality is a substantial improvement compared to any of the previous assemblies, with just 175 scaffolds and having an N50 >135Mb and 756 contigs with N50 >29Mb, resulting in a contiguity similar to the human or mouse reference assemblies. The assembly has been submitted to the International Nucleotide Sequence Database Consortium (INSDC), and the initial GenBank record for it is now available at https://www.ncbi.nlm.nih.gov/assembly/GCA_015227675.1. Genome annotation, i.e. the assignment of gene positions and prediction of new genes and other genomic elements, will be generated by both NCBI and Ensembl.
We are also pleased to announce that Rat and the mRatBN7 assembly have been accepted into the Genome Reference Consortium (GRC) and the RGD has been approved to represent the rat research community and participate in the ongoing work of curating the assembly. RGD will work closely with curators from the GRC, with the International Rat Omics Consortium (IROC), a grassroots community of rat genomics researchers, and the rat research community to identify any candidate regions for focused genome curation. Stay tuned for the appearance of rat on the GRC website!
Monday, November 30, 2020
Wednesday, July 22, 2020
GRCm39: the new mouse reference genome assembly
The GRC is pleased to announce the release of GRCm39 (GCA_000001635.9), the latest version of the mouse reference genome assembly.
GRCm39 is the first coordinate-changing update to the mouse reference since the 2012 release of GRCm38. More than 400 reported issues were resolved in the production of the new assembly, which also incorporates the sequence edits released as scaffolds in the six GRCm38 patch releases.
The new reference assembly exhibits substantial improvements in contiguity. As shown in Fig 1, the scaffold N50 has increased by 95% to 106.1 Mb in GRCm39, and 1.9 Mb of non-N bases were added to the assembly. The gap count has been nearly cut in half, with the total gap length reduced by 4.5 Mb. The decrease in gap length reflects in part the use of optical map data to size the remaining gaps wherever possible, replacing many of the default 50 kb gaps found in GRCm38. Sequences used for gap closures included clones, GRC-constructed contigs, as well as contigs from the C57BL/6J long-read based assembly ASM377452v2.
 |
| Figure 1: GRCm39 Assembly Statistics |
As in prior assembly versions, the GRCm39 chromosome sequences continue to represent the C57BL/6J strain. However, the alternate loci scaffolds that provided additional strain representations for highly variant genomic regions in GRCm38 and MGSCv37, have been removed from the assembly. The relatively low usage of these scaffolds, coupled with a growing number of high quality strain-specific genome assemblies available in public sequence databases, such as those generated by the Mouse Genomes Project, has reduced the need for the inclusion of these sequences in the reference genome assembly. Although no longer affiliated with the reference assembly, these sequences remain available in the INSDC databases (GenBank/ENA/DDBJ).
The new reference assembly will be annotated by GENCODE and RefSeq in the coming months. An in-depth transcript alignment analysis of a pre-release version of the GRCm39 assembly, presented at the 2019 IMGC meeting, demonstrated that there is improved representation for more than 50 genes. A list of these genes is provided in our earlier blog post. The GRC will provide a complete list of genes improved in GRCm39 as the annotation effort progresses.
Notable curation activities represented in the new assembly, but not in previous patch releases, include the targeted update of more than 1,500 individual bases at which the GRCm38 allele representation was erroneous or an unsupported C57BL/6J variant, a substantial retiling of the chr X pseudo-autosomal region (PAR) that provides representation for several genes missing from GRCm38 (Fig 2), removal of a false triplication involving the Duxbl locus, and correction of a 16 Mb inversion at the proximal end of chromosome 14.
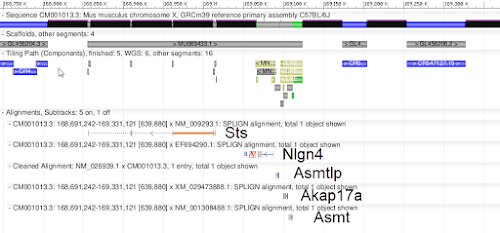 |
| Figure 2. Genes in GRCm39 chr X PAR |
The GRC wishes to thank the many members of the mouse community who have reported assembly issues, and contributed their time, expertise, and data to assist in curation efforts. Updates to the GRC website will be made to reflect the new assembly. With the release of GRCm39, the GRC's curation of the mouse genome reference assembly will be limited to the resolution of community reported problems. We encourage you to contact the GRC for additional information on the curation of assembly regions of interest. You can also subscribe to grc-announce email list to receive email notification for all GRC assembly updates.
Monday, June 8, 2020
ZFIN and the GRC: Supporting the zebrafish reference genome assembly
ZFIN is a member of The Genome Reference Consortium (GRC), an international collaboration consisting of NCBI, the Wellcome Trust Sanger Institute, the McDonnell Genome Institute at Washington University, the European Bioinformatics Institute (EBI) and ZFIN. This consortium is tasked with ensuring that the reference assemblies for human, mouse, zebrafish and chicken are updated and improved through new data and analysis from genome centers and the research community.
The zebrafish-specific GRC webpage (Fig. 1) provides an overview of the zebrafish genome, including an ideogram of the latest zebrafish assembly (GRCz11) that highlights the location of alternate loci scaffolds, downloadable files for the current public assemby and the tiling path files reflecting the latest assembly edits, as well as links to genome assembly data and genome regions under review. Zebrafish genome issues (Fig. 2), such as sequencing errors, gaps, and path problems, can be browsed at the chromosome level, filtered by problem type or status or searched by gene, location, clone name or accession.
If you come across what you suspect is a problem in the build in the course of your research, visit the GRC website to search the list of genome issues and if it has yet to be reported, select the "Report an Issue" tab in the header (Fig. 3) to report information about the potential problem in the build. Be as complete as possible and provide location, flanking sequences and a description of the issue. Genome annotators will evaluate the region, determine if an update to the genome is needed and submit data to create a new tiling path to improve the build with an update or "patch".
We welcome your feedback!
 |
| Figure 1 |
 |
| Figure 2 |
 |
| Figure 3 |
Subscribe to:
Posts (Atom)
